Rapid critical care diagnostics using label-free imaging cytometry
Remoscope analyzes millions of cells from fresh whole blood within minutes without fixation, staining, or any sample preparation. The remo-ID model learns to collect features across hundreds of images, that are predictive of the patient's physician-adjudicated disease state. We trained these models to distinguish sepsis-positive patients from other critically ill patients in the Early Acute Renal and Lung Injury cohort at the University of California San Francisco (EARLI), as well as from a pool of healthy blood donors. The model learns which features are relevant without being told what to look for. When tuned for sepsis rule-out, the model catches 100% of sepsis-positive cases and still achieves 94% specificity.
Sepsis
The red hue shows class-activation — where remo-ID looks when making a call. In these sepsis-positive cases, attention concentrates on abnormal and activated white blood cells. The images shown here are among the highest-confidence positive calls from a few patient samples.
P. falciparum (malaria)
We also trained this type of model to identify malaria (Plasmodium falciparum), which we previously studied using an object detection model called YOGO1. So far, with very little data, the remo-ID attention concentrates on infected red blood cells and overlaps very well with YOGO results. This strengthens our hypothesis that remo-ID is finding disease-predictive features and that extension of the method to many other diseases is possible.
Device and Model
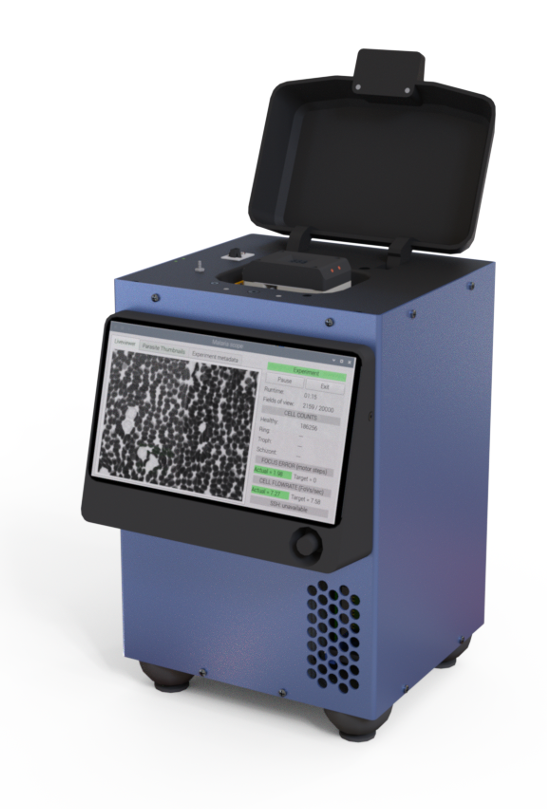

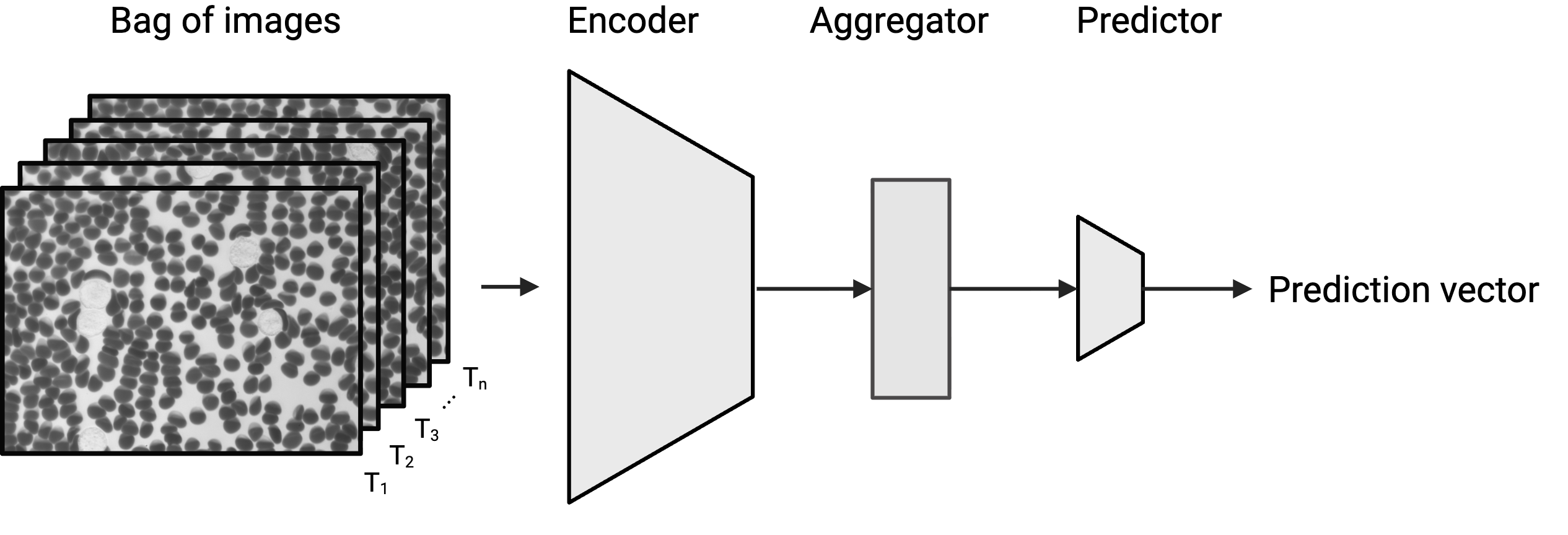
Remoscope is a label-free imaging cytometer that cuts all sample preparation and user training out of the workflow. Fresh whole blood flows through a disposable ultrathin flow cell while millions of cells are imaged in minutes, with on-board compute running end-to-end automation and inference. Here we are able to accurately predict sepsis from just over a hundred images, which can be acquired in seconds.
Quantitative Results
On a small dataset, we were able to train and validate remo-ID models that can distinguish sepsis from other critically ill patients and healthy donors with very high accuracy.Confusion Matrix
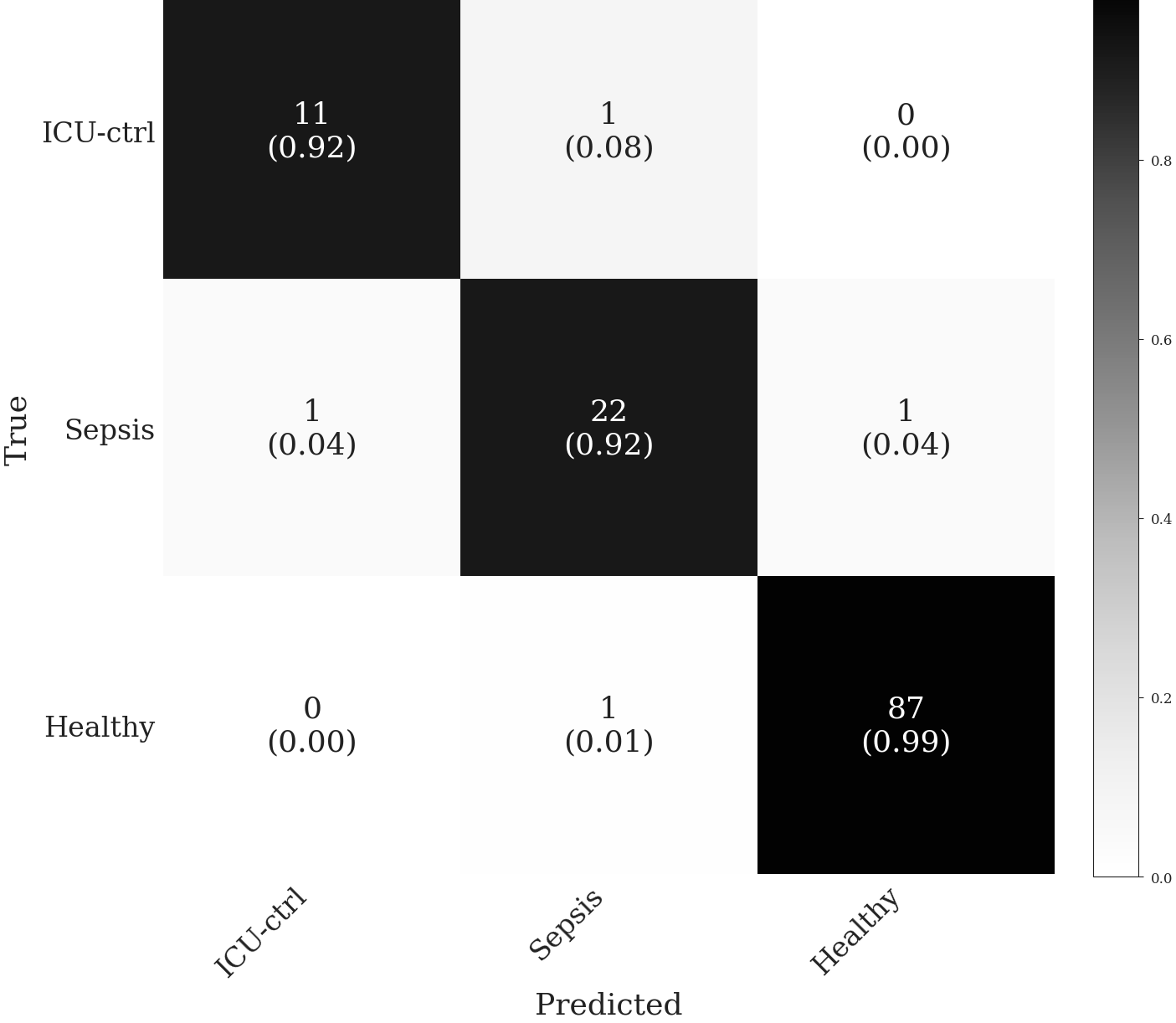

| Class | N | F1 | Sens. | Spec. |
|---|---|---|---|---|
| Sepsis+ | 24 | 0.917 | 0.917 | 0.980 |
| ICU-ctrl | 12 | 0.917 | 0.917 | 0.991 |
| Healthy | 88 | 0.989 | 0.989 | 0.972 |
| Macro | 124 | 0.941 | 0.941 | 0.981 |
ROC Curves (One-vs-Rest)
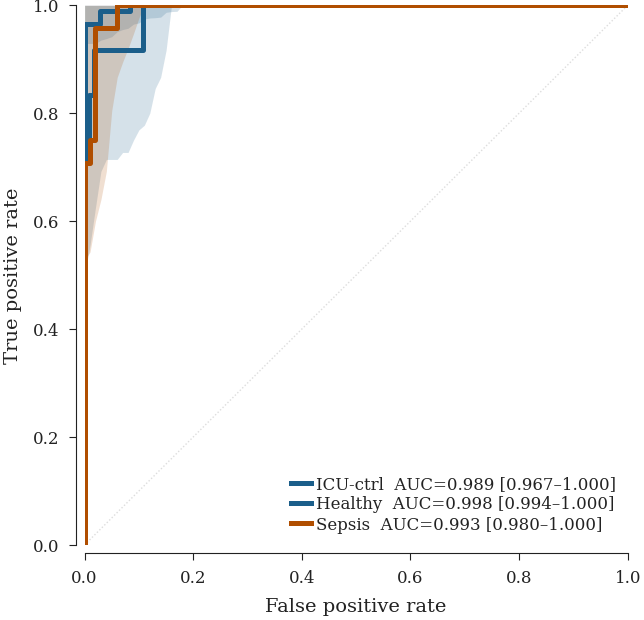
| Comparator | Sens. | Spec. |
|---|---|---|
| vs ICU-ctrl | 100% | 75.0% |
| vs all | 100% | 94.0% |
All three classes are strongly separable, with one-vs-rest AUCs from 0.989 to 0.998. Per-class F1 reaches 0.917 for sepsis and ICU-control and 0.989 for healthy, with a macro F1 of 0.941. Tuned for sepsis rule-out, the model catches every sepsis-positive case at 94% specificity against the combined comparator.
remo-Map: Discovery of cell morphology fingerprints
With remo-Map, we analyze millions of cells and send their images through a powerful embedding model to extract their features and separate them by cell type and state.2 As individuals, cells can be distinguished by type and state. As an ensemble, we can discover patient-level patterns occuring due to meaningful shifts in the distribution of cells that correlates well with their state of health.Here we plot millions of cells from the three classes (healthy, sick-icu, and sepsis), projected onto a UMAP via an unsupervised cell embedding. Click to switch back and forth between segmented cell images, and their average densities in log-scaled blue to orange. In sepsis, white blood cells shift darker and more granulated toward a state of activation.
The Remoscope, equipped with remo-Map and remo-ID, has allowed us to discover disease-predictive features in fresh blood samples that are highly amenable to point of care decision-making. This work is part of an ongoing research study, is subject to change as more results emerge, and will be submitted for peer review on completion along with all appropriate documentation and IRB approval for research on human subjects.
References
- Lebel, P. M. et al. Remoscope: a label-free imaging cytometer for malaria diagnostics. Trans. R. Soc. Trop. Med. Hyg. 119, 1100–1111 (2025). https://doi.org/10.1093/trstmh/traf070
- Bratvold, N. et al. Manuscript in preparation (2026).